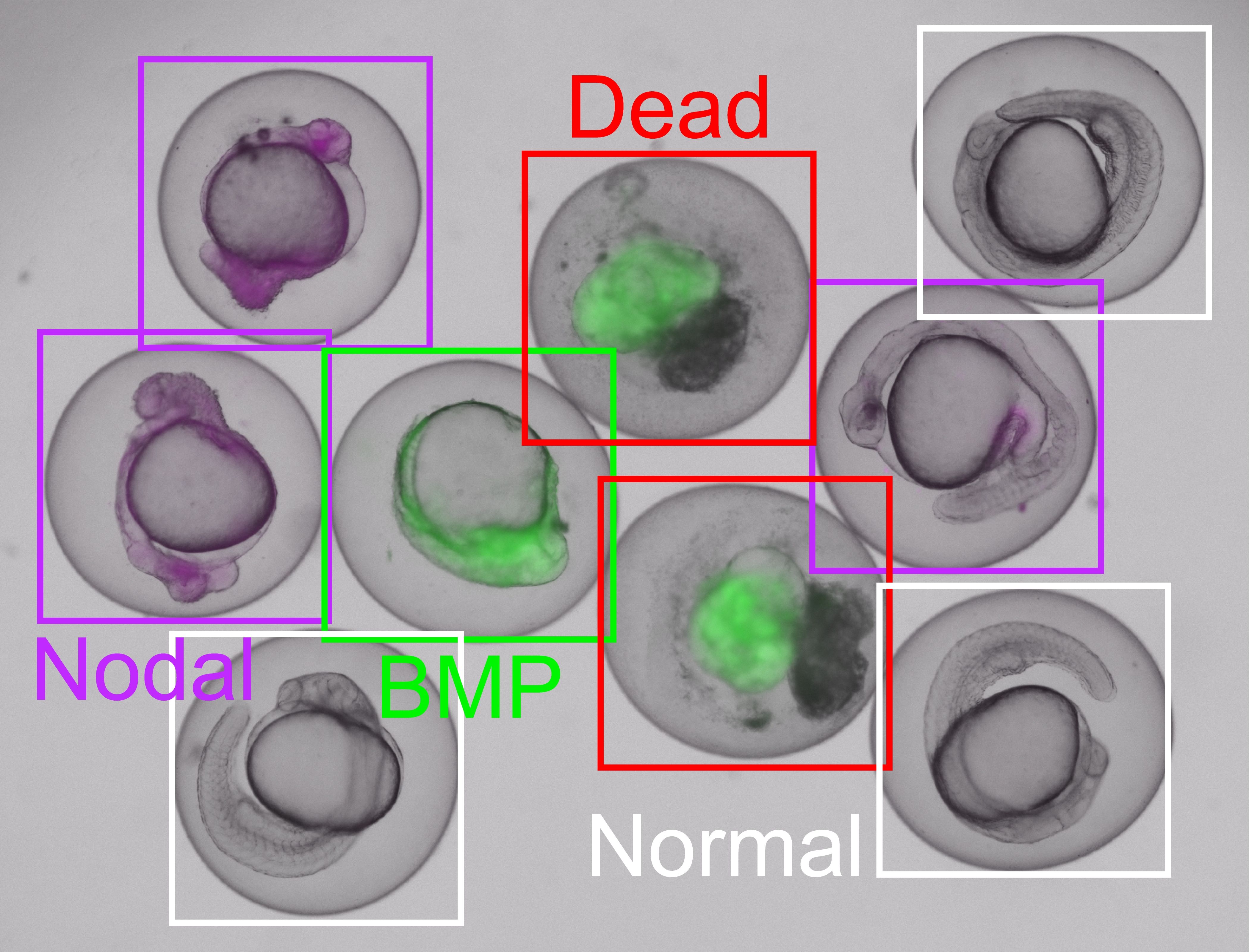
Automated detection of embryonic developmental defects
Researchers at the University of Konstanz publish image analysis software that automatically detects and classifies defects of animal development. Thanks to artificial intelligence, "EmbryoNet" outperforms human experts in terms of speed, accuracy and sensitivity.
Complex multicellular organisms can only emerge from fertilized eggs because embryonic development is biologically precisely regulated. Cellular communication through signaling pathways plays a crucial role in this context. If the activities of the signaling pathways are disturbed, the embryo will show characteristic developmental defects.
In a new study published in the journal Nature Methods, researchers led by Patrick Müller, professor of developmental biology at the University of Konstanz, now present their free software EmbryoNet. The automated image analysis software detects and classifies defects that occur during the development of fish embryos. The classification can then be used to infer which signaling pathway was disturbed in these embryos. In high-throughput applications, the speed and accuracy of the software make it possible to investigate, for example, the mechanisms of how drugs work.
Artificial intelligence as a key component
Up to now, experts were required to microscopically inspect a large number of embryos in order to identify the underlying signaling mechanisms on the basis of visible developmental defects. This time-consuming method is tedious and also prone to differing, partly subjective assessments due to a lack of standardization.
"With EmbryoNet, we are therefore taking a machine learning-based approach, where a neural network trained with over 2 million representative images of zebrafish embryos does the objective classification," reports Matvey Safroshkin, one of the programmers of EmbryoNet along with Hernán Morales-Naverrete. In addition to the image data to be classified, EmbryoNet also takes into account the temporal information on embryonic development and the link between a developmental defect and the corresponding signaling pathway.
More effective than humans
The scientists tested the performance of their software in direct comparison with humans. The task: match previously unclassified images of zebrafish embryos to possible developmental defects. Not only experienced experts in the field of developmental biology competed with EmbryoNet, but also groups of students as part of an undergraduate practical course.
"The students' data were included in our study and are a nice demonstration of how current research and university teaching can benefit from each other", Müller says. The study results show that EmbryoNet can reliably identify different signaling mutants in zebrafish. Moreover, the software was much faster and even more sensitive than its human counterparts – including the experts.
Open-source and adaptable
The researchers also demonstrated that EmbryoNet can be applied not only to zebrafish – a popular model in developmental biology – but also to other vertebrate species. "With relatively little effort, we were able to retrain EmbryoNet to classify other species that evolutionarily separated from zebrafish hundreds of millions of years ago", explains Daniel Čapek, a developmental biologist and one of the study authors. Thus, the open-source software, which is freely usable and modifiable, has the potential to accelerate the characterization of developmental mutants in diverse species.
Key facts:
- Original publication: Čapek, Safroshkin, Morales-Navarrete et al. (2023) EmbryoNet: Using deep learning to link embryonic phenotypes to signaling pathways. Nature Methods; doi: 10.1038/s41592-023-01873-4
- Researchers from Konstanz and Tübingen present EmbryoNet: free software for automated classification of developmental mutants using artificial intelligence
- Scientific contact: Professor Patrick Müller (phone: +49 7531 88 5544; email: patrick.mueller@uni-konstanz.de)
- Funding: European Research Council (ERC) under the European Union’s research and innovation funding programme "Horizon 2020", German Research Foundation (DFG) within the Excellence Strategy, EMBO Young Investigator Programme, Max Planck Society, Medical Faculty of the University of Tübingen and Austrian Science Fund (FWF)
- Open science: EmbryoNet is provided as open-source software along with training and test data for the model presented in the publication on the associated website for free use
